Fast co-evolution of anti-silencing systems shapes the invasiveness of Mu-like DNA transposons in eudicots
Taku Sasaki, Kyudo Ro, Erwann Caillieux, Riku Manabe, Grégoire Bohl-Viallefond, Pierre Baduel, Vincent Colot, Tetsuji Kakutani, Leandro Quadrana
Abstract
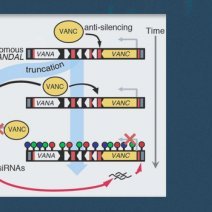
Transposable elements (TEs) constitute a major threat to genome stability and are therefore typically silenced by epigenetic mechanisms. In response, some TEs have evolved counteracting systems to suppress epigenetic silencing. In the model plant Arabidopsis thaliana, two such anti-silencing systems have been identified and found to be mediated by the VANC DNA-binding proteins encoded by VANDAL transposons. Here, we show that anti-silencing systems have rapidly diversified since their origin in eudicots by gaining and losing VANC-containing domains, such as DUF1985, DUF287, and Ulp1, as well as target sequence motifs. We further demonstrate that these motifs determine anti-silencing specificity by sequence, density, and helical periodicity. Moreover, such rapid diversification yielded at least 10 distinct VANC-induced anti-silencing systems in Arabidopsis. Strikingly, anti-silencing of non-autonomous VANDALs, which can act as reservoirs of 24-nt small RNAs, is critical to prevent the demise of cognate autonomous TEs and to ensure their propagation. Our findings illustrate how complex co-evolutionary dynamics between TEs and host suppression pathways have shaped the emergence of new epigenetic control mechanisms.
The EMBO Journal (2022)e110070. doi 10.15252/embj.2021110070