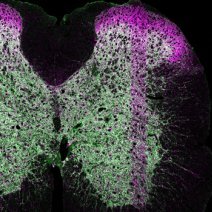
Identification of a stereotypic molecular arrangement of endogenous glycine receptors at spinal cord synapses
S A Maynard, P Rostaing, N Schaefer, O Gemin, A Candat, A Dumoulin, C Villmann, A Triller, C G Specht
Abstract
Precise quantitative information about the molecular architecture of synapses is essential to understanding the functional specificity and downstream signaling processes at specific populations of synapses. Glycine receptors (GlyRs) are the primary fast inhibitory neurotransmitter receptors in the spinal cord and brainstem. These inhibitory glycinergic networks crucially regulate motor and sensory processes. Thus far, the nanoscale organization of GlyRs underlying the different network specificities has not been defined. Here, we have quantitatively characterized the molecular arrangement and ultra-structure of glycinergic synapses in spinal cord tissue using quantitative super-resolution correlative light and electron microscopy. We show that endogenous GlyRs exhibit equal receptor-scaffold occupancy and constant packing densities of about 2000 GlyRs µm-2 at synapses across the spinal cord and throughout adulthood, even though ventral horn synapses have twice the total copy numbers, larger postsynaptic domains, and more convoluted morphologies than dorsal horn synapses. We demonstrate that this stereotypic molecular arrangement is maintained at glycinergic synapses in the oscillator mouse model of the neuromotor disease hyperekplexia despite a decrease in synapse size, indicating that the molecular organization of GlyRs is preserved in this hypomorph. We thus conclude that the morphology and size of inhibitory postsynaptic specializations rather than differences in GlyR packing determine the postsynaptic strength of glycinergic neurotransmission in motor and sensory spinal cord networks.
Keywords
correlative light and electron microscopy ; electron microscopy ; glycine receptor ; hyperekplexia ; mouse ; neuroscience ; oscillator mouse model ; single molecule localization microscopy ; spinal cord.
© 2021, Maynard et al.
Elife. 2021 Dec 8 ;10:e74441. doi : 10.7554/eLife.74441