The downloaded FASTQ file is available in the data folder (SRR576938.fastq)
Introduction
GoalThe aim is to :
- have an understanding of the nature of ChIP-Seq data
- perform a complete analysis workflow including quality check (QC), read mapping, visualization in a genome browser and peak-calling. Use command line and open source software for each each step of the workflow and feel the complexity of the task
- have an overview of possible downstream analyses
- perform a motif analysis with online web programs
This training gives an introduction to ChIP-seq data analysis, covering the processing steps starting from the reads to the peaks. Among all possible downstream analyses, the practical aspect will focus on motif analyses. A particular emphasis will be put on deciding which downstream analyses to perform depending on the biological question. This training does not cover all methods available today. It does not aim at bringing users to a professional NGS analyst level but provides enough information to allow biologists understand what DNA sequencing practically is and to communicate with NGS experts for more in-depth needs.
Dataset descriptionFor this training, we will use a dataset produced by Myers et al (see on the right of the page for details on the publication) involved in the regulation of gene expression under anaerobic conditions in bacteria. We will focus on one factor: FNR.
Downloading ChIP-seq reads from NCBI
Related VIB training: Downloading NGS data from NCBI
1 - Obtaining an identifier for a chosen dataset
Within an article of interest, search for a sentence mentioning the deposition of the data in a database. Here, the following sentence can be found at the end of the Materials and Methods section:
"All genome-wide data from this publication have been deposited in NCBI’s Gene Expression Omnibus (GSE41195)."
We will thus use the GSE41195 identifier to retrieve the dataset from the NCBI GEO (Gene Expression Omnibus) database.
NGS datasets are (usually) made freely accessible for other scientists, by depositing these datasets into specialized databanks. Sequence Read Archive (SRA) located in USA hosted by NCBI, and its european equivalent European Nucleotide Archive (ENA) located in England hosted by EBI both contains raw reads.
Functional genomic datasets (transcriptomics, genome-wide binding such as ChIP-seq,...) are deposited in the databases Gene Expression Omnibus (GEO) or its European equivalent ArrayExpress.
2 - Accessing GSE41195 from GEO
- The GEO database hosts processed data files and many details related to the experiments. The SRA (Sequence Read Archive) stores the actual raw sequence data.
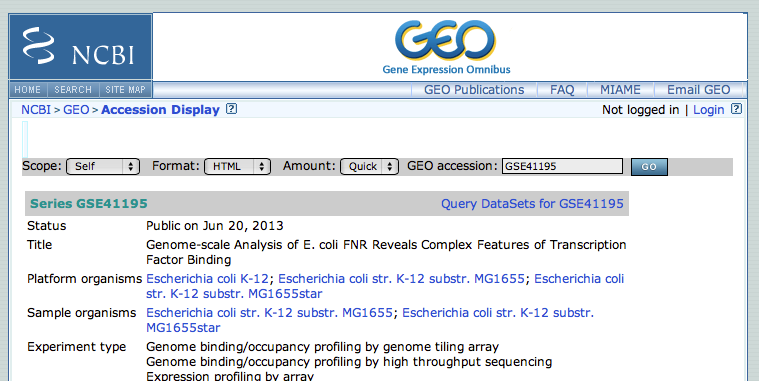
- Search in google GSE41195. Click on the first link to directly access the correct page on the GEO database.
- This GEO entry is a mixture of expression analysis and chip-seq. At the bottom of the page, click on the subseries related to the chip-seq datasets. (this subseries has its own identifier:GSE41187)
- From this page, we will focus on the experiment FNR IP ChIP-seq Anaerobic A. At the bottom of the page, click on the link "GSM1010219 - FNR IP ChIP-seq Anaerobic A"
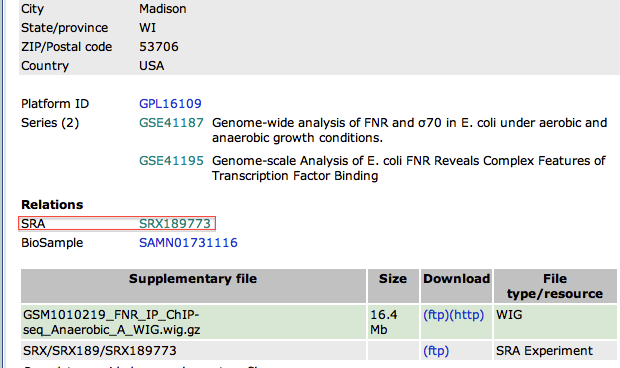
- In the new page, go to the bottom to find the SRA identifier. This is the identifier of the raw dataset stored in the SRA database.
- Copy the identifier SRX189773 (do not click on the link that would take you to the SRA database, see below why)

3 - Downloading FASTQ file from the ENA database
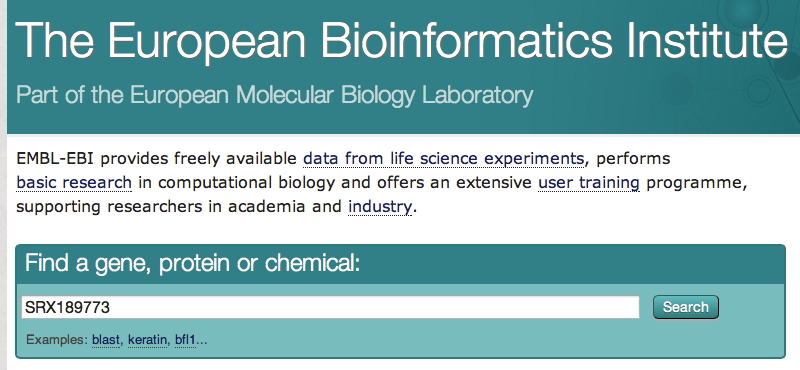
- Go to the EBI website. Paste your SRA identifier (SRX189773) and click on the button "search"
- Click on the first result. On the next page, there is a link to the FASTQ file. For efficiency, this file has already been downloaded and is available in the "data" folder (SRR576933.fastq)
Quality control of the reads and statistics
Related VIB training: Quality control of NGS data
1 - Getting the FASTQC report
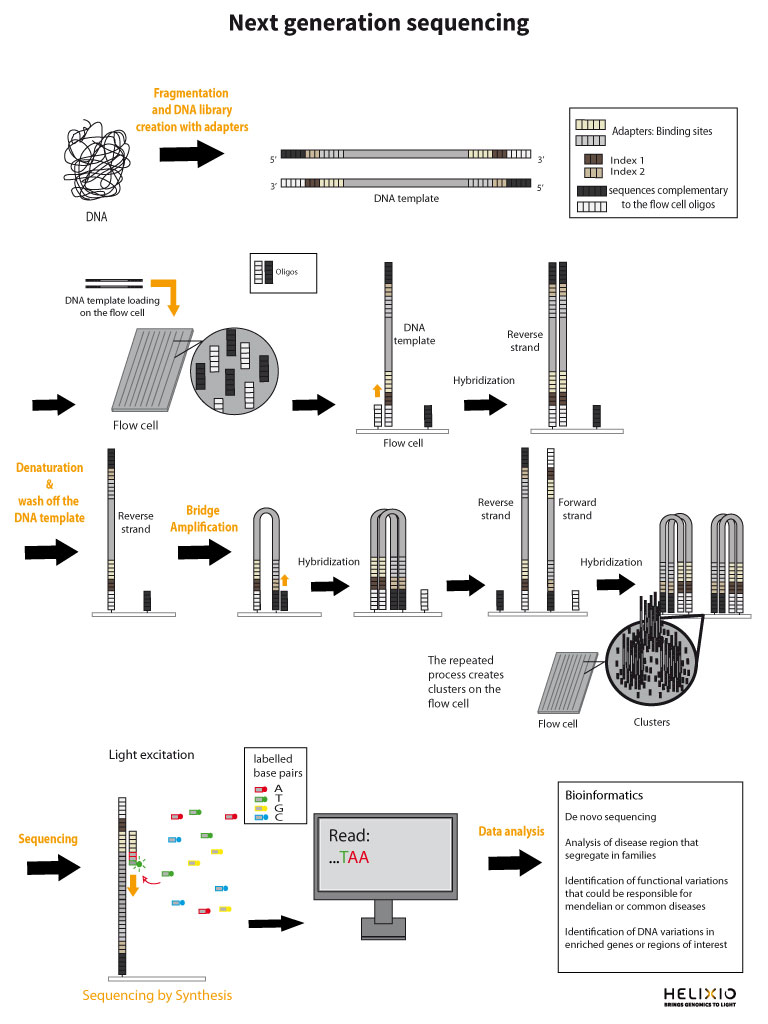
Before you analyze the data, it is crucial to check the quality of the data. We will use the standard tool for checking the quality of data generated on the Illumina platform: FASTQC. Now is the time to gather your courage and face the terminal. Don't worry, it's not as scary as it seems ! Below, the "$" sign is to indicate what you must type in the command line, but don't type the "$".
- Open the terminal and go to the course directory.
- First check the help page of the program to see its usage and parameters.
- Launch the FASTQC program on the experiment file (SRR576933.fastq)
- Wait until the analysis is finished. Check the files output by the program.
$ ls SRR576933_fastqc SRR576933_fastqc.zip SRR576933.fastq
- Access the result
$ cd SRR576933_fastqc $ ls fastqc_data.txt fastqc_report.html Icons Images summary.txt
- Open the HTML file fastqc_report.html in Firefox.
- Now do the same with the control (Input) dataset (SRR576938.fastq).
- Go to the NCBI Genome website, and search for the organism Escherichia coli
- Click on the Escherichia coli str. K-12 substr. MG1655 to access statistics on this genome.
$ cd /home/bits/NGS/ChIPSeq
fastqc --help
$ fastqc SRR576933.fastq
How many reads are present in the file ?
What is the read length ? Is the overall quality good ?
Are there any concerns raised by the report ? If so, can you tell where the problem might come from ?
There are 3 603 544 reads of 36bp. The overall quality is good, although it drops at the last position, which is usual with Illumina sequencing, so this feature is not raising hard concerns. There are several "red lights" in the report. In particular, the per sequence GC content and the duplication level are problematic. If you check the "overrepresented sequences", you'll notice a high percentage of adapters (29% !). Ideally, we would remove these adapters (=trim) the reads, and then re-run FASTQC. In practice, we often skip this step, as these reads will anyway not be mapped. Warning: this will affect the future calculated "% of mapped read" !!
There are 6 717 074 reads of 36bp. The overall quality is good, although it drops at the last position, which is usual with Illumina sequencing, so this feature is not raising hard concerns. The duplication level is high, which could be due to the high coverage. We will check this later, by visualizing the dataset in a genome browser.
2 - Organism length
Knowing your organism size is important to evaluate if your dataset has sufficient coverage to continue your analyses. For the human genome (3 Gb), we usually aim at least 10 Million reads.
Do both FASTQ files contain enough reads for a proper analysis ?
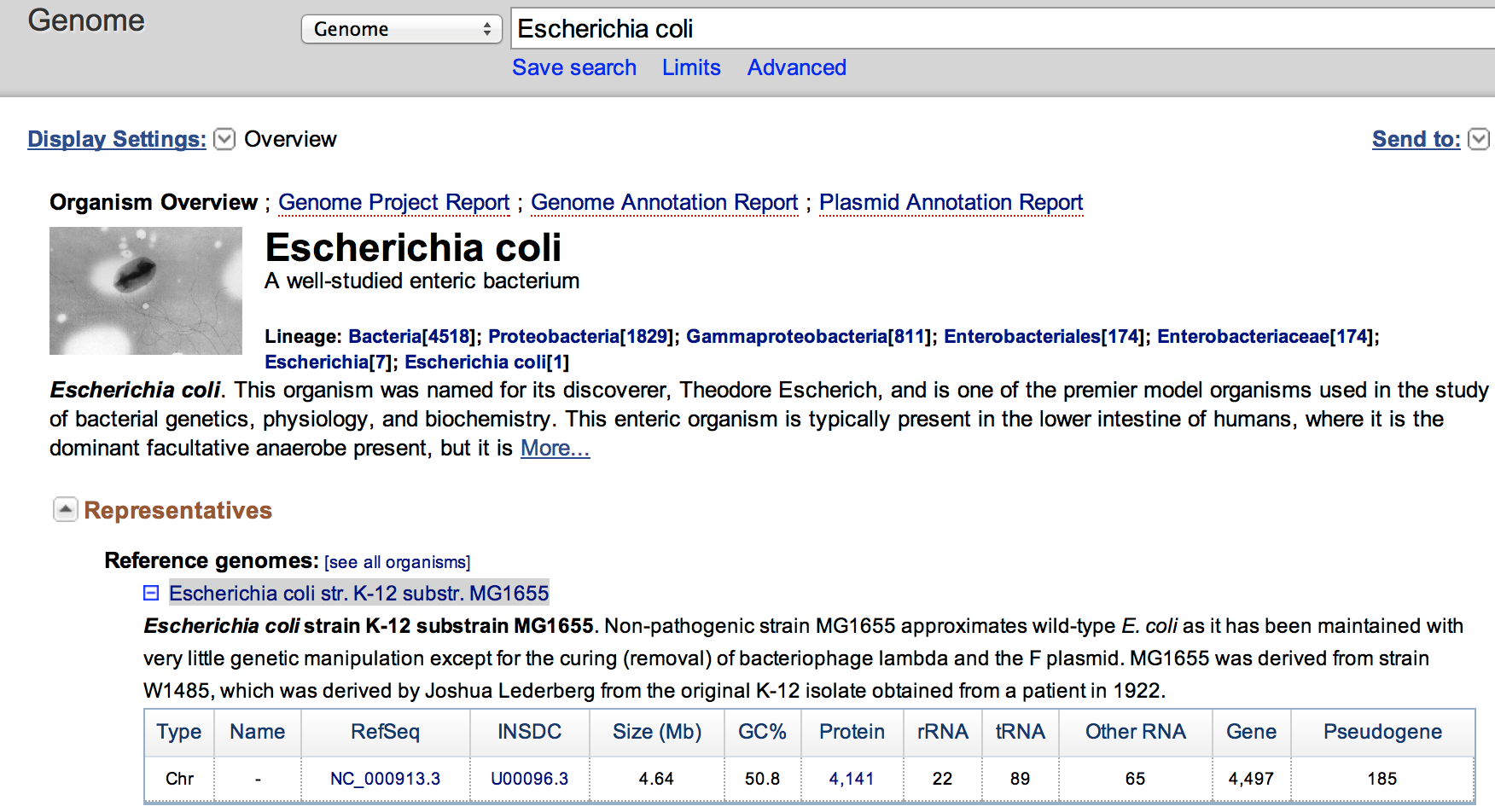
The genome is 4.64 Mbase. The files have respectively 3.6 M reads and 6.7 M reads. As we aim for 10 M for 3 Gb when working with human, the dataset here should enough reads for proper analysis.
Mapping the reads with Bowtie
Related VIB training: Mapping of NGS data
1 - Choosing a mapping program
There are multiple programs to perform the mapping step. For reads produced by an Illumina machine for ChIP-seq, the currently "standard" programs are BWA and Bowtie (versions 1 and 2), and STAR is getting popular. We will use Bowtie version 1.1.1 for this exercice, as this program remains effective for short reads (< 50bp).
2 - Prepare the index file
- Try out bowtie
- bowtie needs the reference genome to align each read on it. This genome needs to be in a specific format (=index) for bowtie to be able to use it. Several pre-built indexes are available to download on the bowtie webpage, but our genome is not there. You will need to make this index file.
- To make the index file, you will need the complete genome, in FASTA format. It has already been downloaded to gain time (Escherichia_coli_K12.fasta in the course folder) (The genome was downloaded from the NCBI). Note that we will not work with the lastest version (NC_000913.3) but the previous one (NC_000913.2), because the available tools for visualization have not been updated yet to the latest version. This will not affect our results.
- Build the index for bowtie
$ bowtie-1.1.1This prints the help of the program. However, this is a bit difficult to read ! If you need to know more about the program, it's easier to directly check the manual on the website (if it's up-to-date !)
$ bowtie-1.1.1-build Escherichia_coli_K12.fasta Escherichia_coli_K12
3 - Mapping the experiment
- Let's see the parameters of bowtie before launching the mapping:
- Escherichia_coli_K12 is the name of our genome index file
- Number of mismatches for SOAP-like alignment policy (-v): to 2, which will allow two mismatches anywhere in the read, when aligning the read to the genome sequence.
- Suppress all alignments for a read if more than n reportable alignments exist (-m): to 1, which will exclude the reads that do not map uniquely to the genome.
- -q indicates the input file is in FASTQ format. SRR576933.fastq is the name of our FASTQ file.
- -3 will trim x base from the end of the read. As our last position is of low quality, we'll trim 1 base.
- -S will output the result in SAM format
- 2> SRR576933.out will output some statistics about the mapping in the file SRR576933.out
- This should take few minutes as we work with a small genome. For the human genome, we would need either more time, or a dedicated server.
$ bowtie-1.1.1 Escherichia_coli_K12 -q SRR576933.fastq -v 2 -m 1 -3 1 -S 2> SRR576933.out > SRR576933.sam
Open the file SRR576933.out, which contains some statistics about the mapping. How many reads were mapped ?
$ less SRR576933.out # reads processed: 3603544 # reads with at least one reported alignment: 2242431 (62.23%) # reads that failed to align: 1258533 (34.92%) # reads with alignments suppressed due to -m: 102580 (2.85%) Reported 2242431 alignments to 1 output stream(s)Type q to go back. 62% of mapped reads is good, especially as we know there were adapter sequences in 29% of the reads.
4 - Mapping the control
- Repeat the steps above (step 3) for the file SRR576938.fastq.
Open the file SRR576938.out. How many reads were mapped ?
$ bowtie-1.1.1 Escherichia_coli_K12 -q SRR576938.fastq -v 2 -m 1 -3 1 -S 2> SRR576938.out > SRR576938.sam
$less SRR576938.out # reads processed: 6717074 # reads with at least one reported alignment: 6393027 (95.18%) # reads that failed to align: 107750 (1.60%) # reads with alignments suppressed due to -m: 216297 (3.22%) Reported 6393027 alignments to 1 output stream(s)95% of mapped reads is very good.
Bonus: checking two ENCODE quality metrics
1 - PhantomPeakQualTools
Warning : this exercice is new and is tested for the first time in a classroom context. Let us know if you encounter any issue.
The SAM format correspond to large text files, that can be compressed ("zipped") into BAM format. The BAM format are usually sorted and indexed for fast access to the data it contains.
- convert the SAM file into a sorted BAM file
$ samtools-0.1.19 view -bS SRR576933.sam > SRR576933.bam
- convert the BAM file into TagAlign format, specific to the program that calculates the quality metrics
- Run phantompeakqualtools
$ samtools-0.1.19 view -F 0x0204 -o - SRR576933.bam | awk 'BEGIN{OFS="\t"}{if (and($2,16) > 0) {print $3,($4-1),($4-1+length($10)),"N","1000","-"} else {print $3,($4-1),($4-1+length($10)),"N","1000","+"} }' | gzip -c > SRR576933_experiment.tagAlign.gz
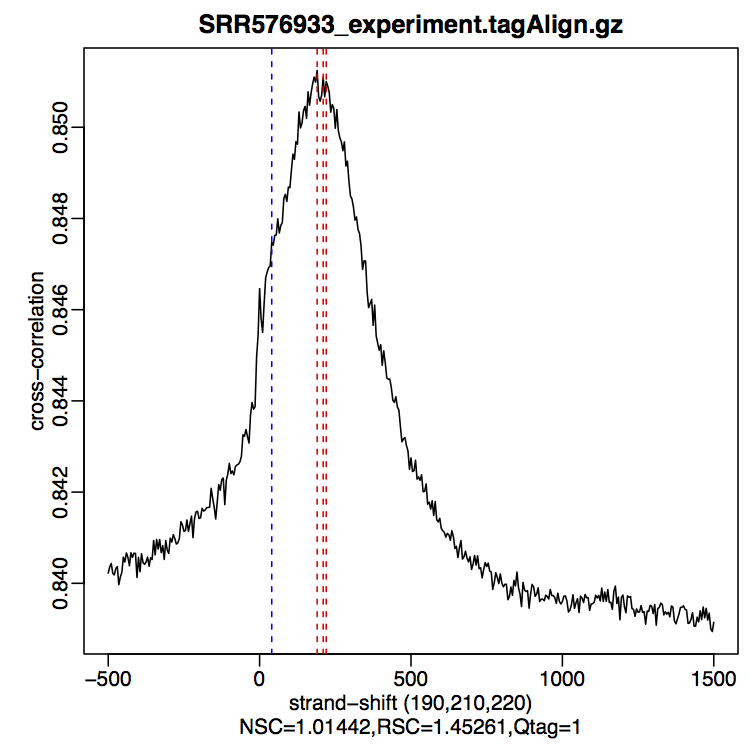
$ Rscript /usr/bin/tools/phantompeakqualtools/run_spp.R -c=SRR576933_experiment.tagAlign.gz -savp -out=SRR576933_experiment_phantompeaks
According to the ENCODE guidelines, NSC >=� 1.05 ; RSC >= �0.8 is recommended. Qtag values range from -2,-1,0,1,2
What is the quality of this dataset ?
Warning: these values are for vertebrate, so in the context of this E. coli dataset, it is difficult to interpret these values correctly. For more details, see Marinov et al, 2014.
Peak calling with MACS
1 - Choosing a peak-calling program
There are multiple programs to perform the peak-calling step. Some are more directed towards histone marks (broad peaks) while others are specific to narrow peaks (transcription factors). Here we will use MACS version 1.4.2 because it's known to produce generally good results, and it is well-maintained by the developper. A new version (MACS2) is being developped, but still in testing phase so we will not use it today.
2 - Calling the peaks
- Try out MACS
- Let's see the parameters of MACS before launching the mapping:
- ChIP-seq tag file (-t) is the name of our experiment (treatment) mapped read file SRR576933.sam
- ChIP-seq control file (-c) is the name of our input (control) mapped read file SRR576938.sam
- --format SAM indicates the input file are in SAM format. Other formats can be specified (BAM,BED...)
- --gsize Effective genome size: this is the size of the genome considered "usable" for peak calling. This value is given by the MACS developpers on their website. It is smaller than the complete genome because many regions are excluded (telomeres, highly repeated regions...). The default value is for human (2700000000.0), so we need to chnage it. As the value for E. coli is not provided, we will take the complete genome size 4639675.
- --name provides a prefix for the output files. We set this to macs14, but it could be any name.
- --bw The bandwidth is the size of the fragment extracted from the gel electrophoresis or expected from sonication. By default, this value is 300bp. Usually, this value is indicated in the Methods section of publications. In the studied publication, a sentence mentions "400bp fragments (FNR libraries)". We thus set this value to 400.
- --keep-dup specifies how MACS should treat the reads that are located at the exact same location (duplicates). The manual specifies that keeping only 1 representative of these "stacks" of reads is giving the best results.
- --bdg --single-profile will output a file in BEDGRAPH format to visualize the peak profiles in a genome browser. There will be one file for the treatment, and one for the control.
- --diag is optional and increases the running time. It tests the saturation of the dataset, and gives an idea of how many peaks are found with subsets of the initial dataset.
- &> MACS.out will output the verbosity (=information) in the file MACS.out
- This should take about 10 minutes, mainly because of the
--diagand the--bdgoptions. Without, the program runs much faster.
$ macs14This prints the help of the program.
$ macs14 -t SRR576933.sam -c SRR576938.sam --format SAM --gsize 4639675 --name "macs14" --bw 400 --keep-dup 1 --bdg --single-profile --diag &> MACS.out
3 - Analyzing the MACS results
$ ls -h MACS.out macs14_MACS_bedGraph macs14_diag.xls macs14_negative_peaks.xls macs14_peaks.bed macs14_peaks.xls macs14_summits.bed
- MACS.out: information on MACS run.
- macs14_diag.xls: diagnostic file. Here, we see that most of the peaks (83%) are found with only 20% of the initial dataset, indicating saturation.
- macs14_negative_peaks.xls: peaks called when inverting the treatment and the control.
- macs14_peaks.bed: peak coordinates in BED format
- macs14_peaks.xls: peak coordinates with more information, to be opened with Excel
- macs14_summits.bed: location of the summit base for each peak (BED format)
$ wc -l macs14_peaks.bed109 peaks were called with these parameters. In the article, the authors found 207 peaks, by integrating the results of 3 peak-calling programs and additional ChIP-chip data.
Visualizing the peaks in a genome browser
Related VIB training: Visualize all results in the Interactive Genome Viewer (IGV)
1 - Choosing a genome browser
There are several options for genome browsers, divided between the local browsers (need to install the program, eg. IGV) and the online web browsers (eg. UCSC genome browser, Ensembl). We often use both types, depending on the aim and the localisation of the data.
If the data are on your computer, to prevent data transfer, it's easier to visualize the data locally (IGV). Note that if you're working on a non-model organism, the local viewer will be the only choice. If the aim is to share the results with your collaborators, view many tracks in the context of many existing annotations, then the online genome browsers are more suitable.
2 - Viewing the peaks in IGV
- Open IGV
- Load the desired genome:
- Unzip the bedgraph files and rename them with the extension .bedgraph, otherwise the file will not be recognized by IGV.
- View the bedgraph files
- View the BED file containing the peak locations
- To see the genes, load the file Escherichia_coli_K_12_MG1655.annotation.fixed.bed
$ igv
(from the fasta file Escherichia_coli_K_12_MG1655.fasta, used to build the bowtie index file)
The loaded genome appears in the top left panel:
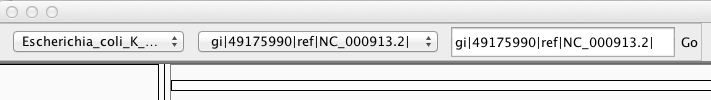
$ gunzip macs14_control_afterfiting_all.bdg.gz
mv macs14_control_afterfiting_all.bdg macs14_control_afterfiting_all.bedgraphDo the same for the treatment file
You might get a warning that the file is big. Simply click on the button continue.
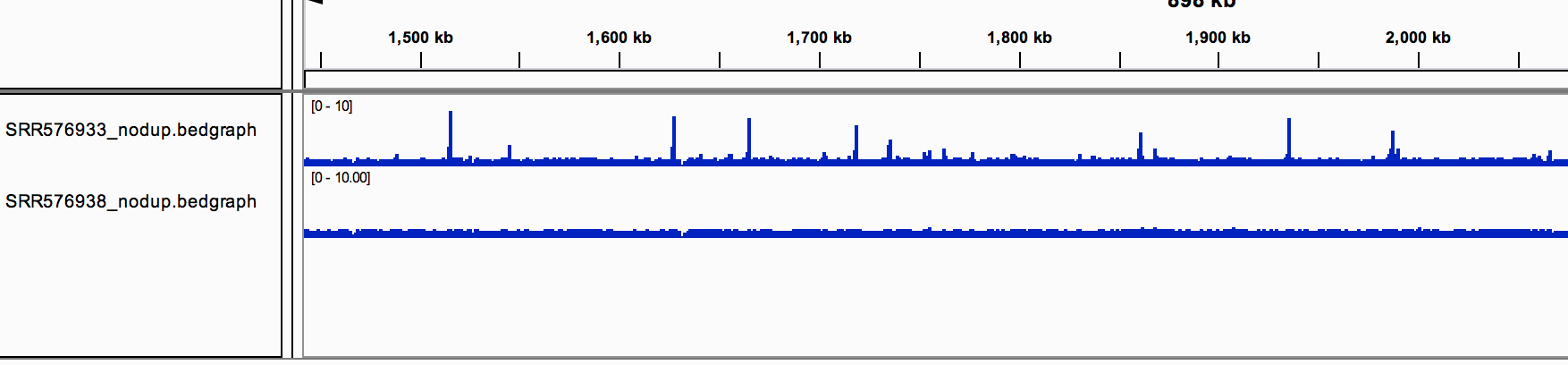
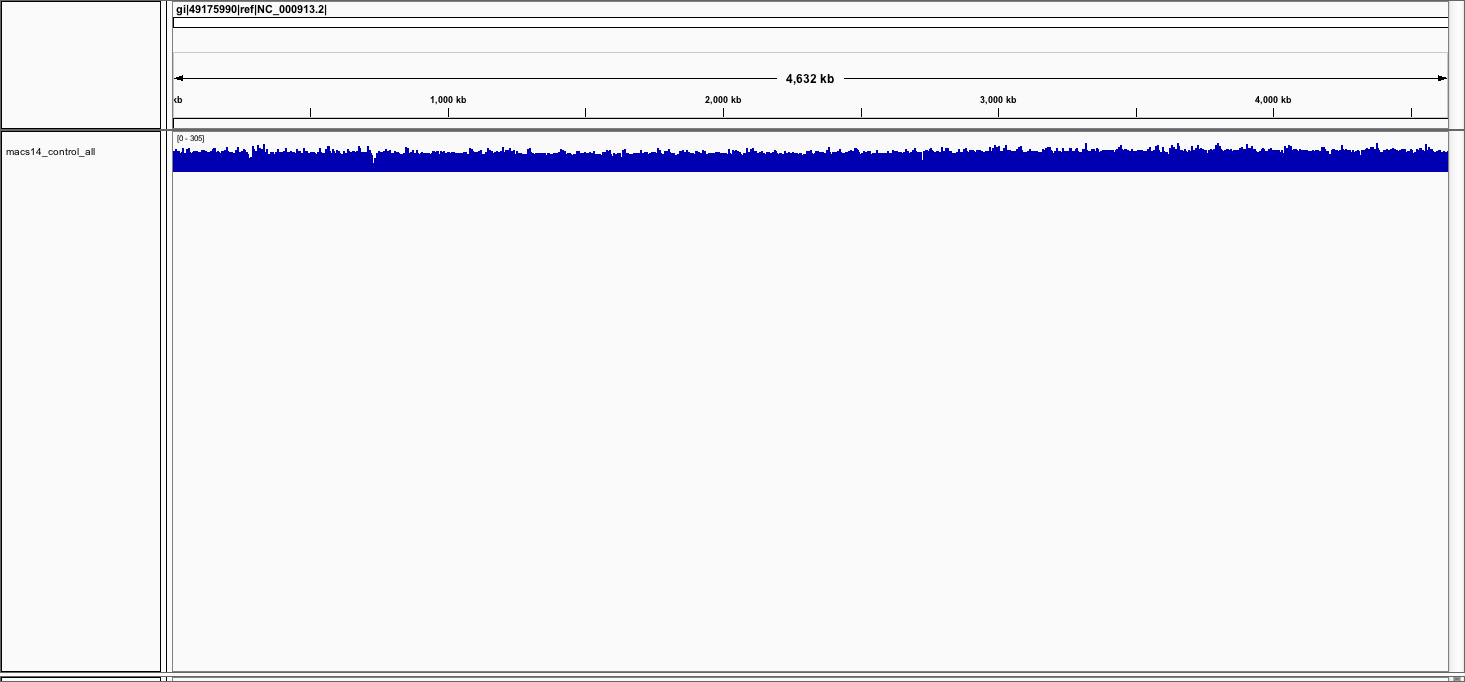 You should see the track (in blue).
You should see the track (in blue).
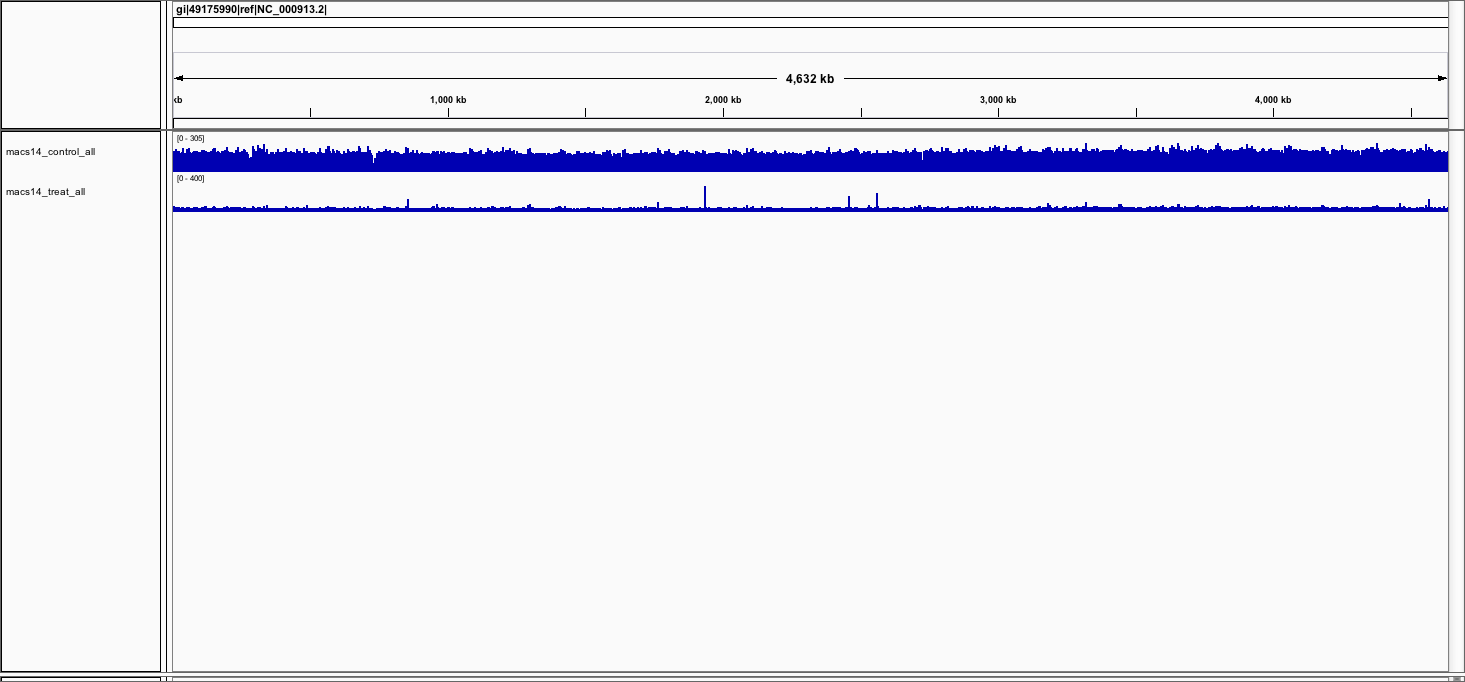
You should now see the 2 tracks (in blue).
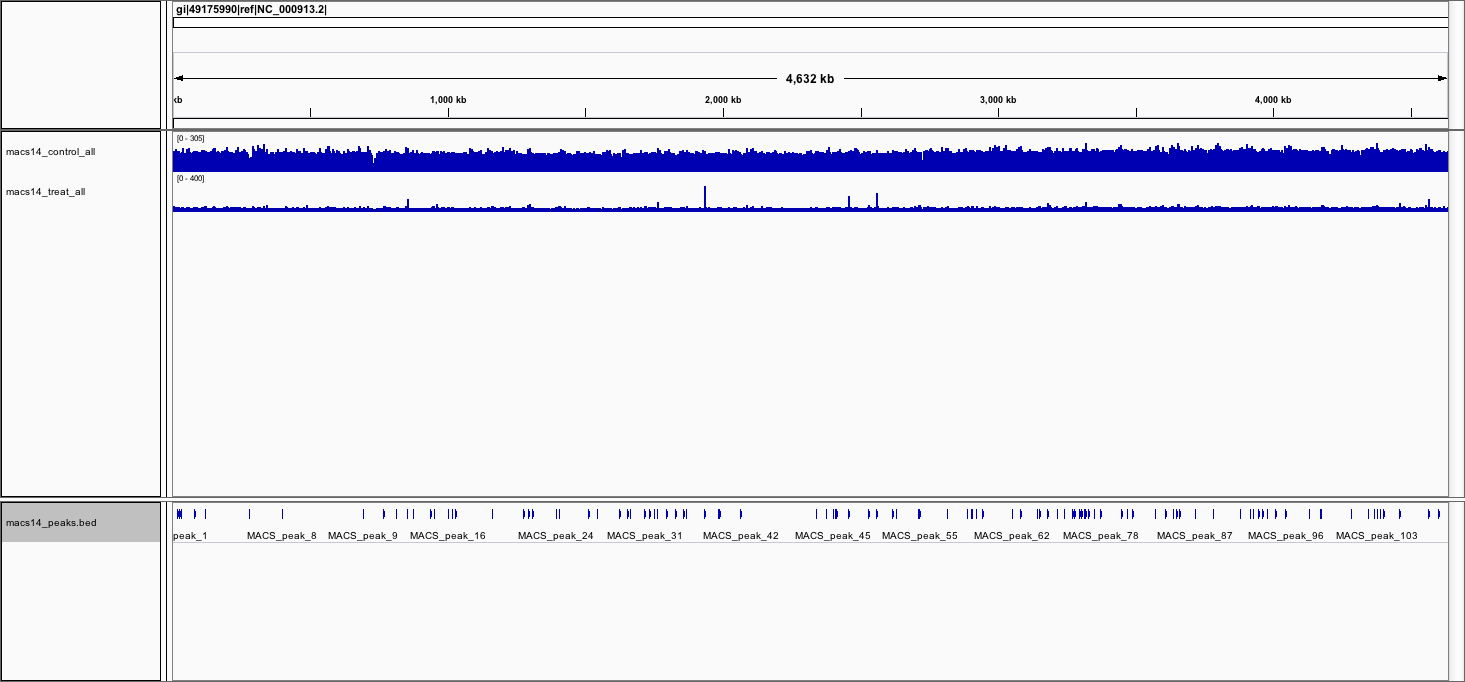
Bonus: vizualisation with deeptools
1 - From SAM to sorted BAM
The SAM format correspond to large text files, that can be compressed ("zipped") into BAM format. The BAM format are usually sorted and indexed for fast access to the data it contains.
- convert the SAM file into a sorted BAM file
$ samtools-0.1.19 view -bS SRR576933.sam | samtools-0.1.19 sort - SRR576933_sorted
- Remove the duplicated reads
- Index the BAM file
$ samtools-0.1.19 rmdup -s SRR576933_sorted.bam SRR576933_sorted_nodup.bamfrom 2242431 uniquely mapped reads, it remains 1142724 reads without duplicates.
$ samtools-0.1.19 index SRR576933_sorted_nodup.bam SRR576933_sorted_nodup.bai
2 - From BAM to scaled Bedgraph files
- bamCoverage from deepTools generates BedGraphs from BAM files
$ bamCoverage --help
- Generate a scaled BedGraph file
$ bamCoverage --bam SRR576933_sorted_nodup.bam --outFileName SRR576933_nodup.bedgraph --outFileFormat bedgraph --normalizeTo1x 4639675
Motif analysis
1 - Retrieve the peak sequences corresponding to the peak coordinate file (BED)
For the motif analysis, you first need to extract the sequences corresponding the the peaks. There are several ways to do this (as usual...). If you work on a UCSC-supported organism, the easiest is to use RSAT fetch-sequences or Galaxy .
Here, we will use Bedtools, as we have the genome of our interest on our computer (Escherichia_coli_K_12_MG1655.fasta).
$ bedtools getfasta -fi Escherichia_coli_K_12_MG1655.fasta -bed macs14_peaks.bed -fo macs14_peaks.fa
$ grep -c '^>' macs14_peaks.fa
2 - Motif discovery with RSAT
- Open a connection to a Regulatory Sequence Analysis Tools server. You can choose between various website mirrors.
- Server at Roscoff (recommended for this training) rsat.sb-roscoff.fr
- Main server (currently in Brussels) www.rsat.eu
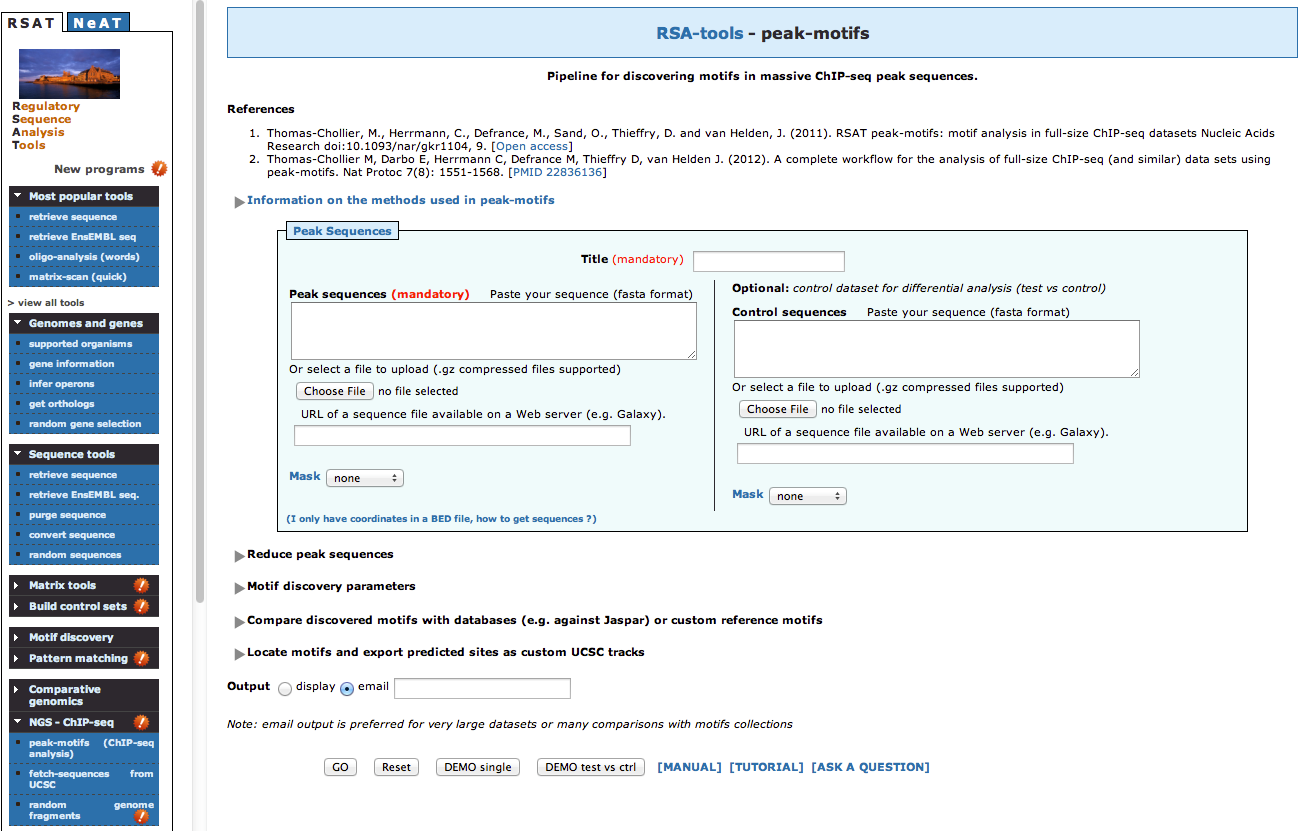
- In the left menu, click on NGS ChIP-seq and then click on peak-motifs. A new page opens, with a form
- The default peak-motifs web form only displays the essential options. There are only two
mandatory parameters.
a. The title box, which you will set as FNR Anaerobic A b. The sequences, that you will upload from your computer, by clicking on the buttonChoose file, and select the file macs14_peaks.fa from your computer. - We could launch the analysis like this, but we will now modify some of the advanced options in order to fine-tune the analysis according
to your data set.
- Open the "Reduce peak sequences" title, and make sure the "Cut peak sequences: +/- " option is set to 0 (we wish to analyse our full dataset)
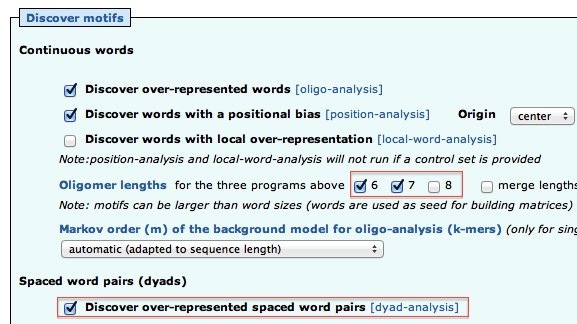
- Open the “Motif Discovery parameters” title, and check the oligomer sizes 6 and 7 (but not 8). Check "Discover over-represented spaced word pairs [dyad-analysis]"
- Under “Compare discovered motifs with databases”, uncheck "JASPAR core vertebrates" and check RegulonDB prokaryotes (2012_05) as the studied organism is the bacteria E. coli.
- You can indicate your email address in order to receive notification of the task submission and completion. This is particularly useful because the full analysis may take some time for very large datasets.
- Click “GO”. As soon as the query has been launched, you should receive an email indicating confirming the task submission, and providing a link to the future result page.
- The Web page also displays a link, You can already click on this link. The report will be progressively updated during the processing of the workflow.
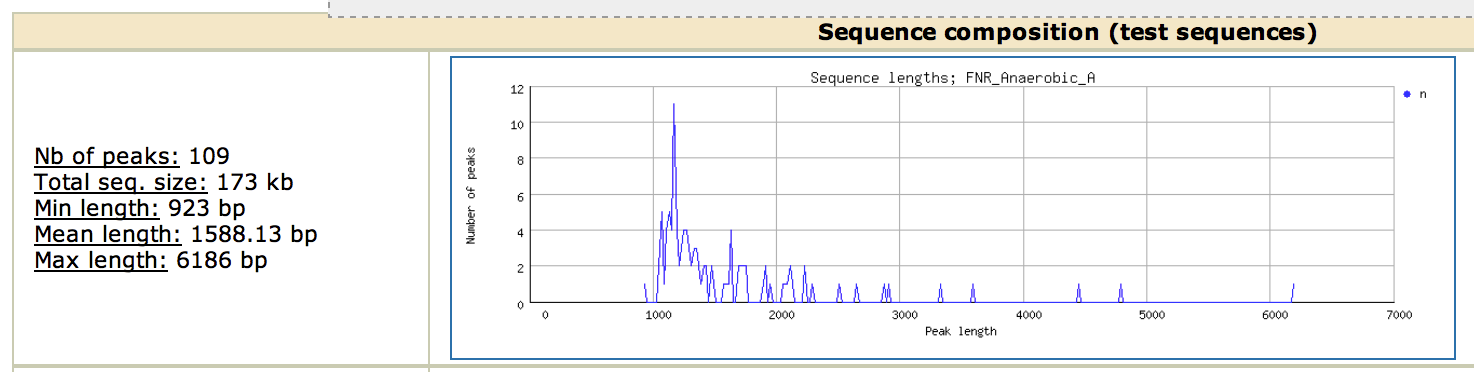
This section is extremely important, as it gives you information about your dataset. Here, we see that our 109 peaks are extremely long : a minimal length of 928bp and a mean of 1.5kb, while the fragment size was 400 bp!! Note that this is a know issue with MACS, that tends to output large peaks. For pedagogic purposes, this step was important to show the importance of peak calling on the motif analysis step. There are different options to overcome this issue: take only the region around the summit of the peaks (often a good solution, see below) or try an alternative peak-calling program (Note that this issue is apparently resolved in the next version of MACS).
3 - Motif discovery with RSAT (short peaks)
- Restrict the dataset to the summit of the peaks +/- 100bp
- Extract the sequences for this BED file
- Run RSAT peak-motifs with the same options, but choosing as input file this new dataset (macs14_summits+-100.fa)
and setting the title box to FNR Anaerobic A summit +/-100bpThis time, do we discover significant motifs ? Are these motifs biologically relevant? In particular, did the program discover motifs related to FNR ?
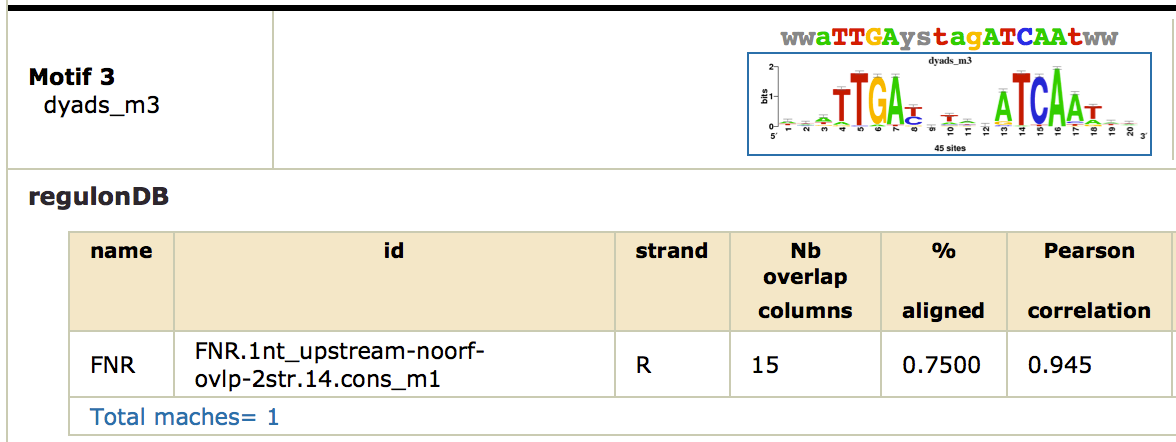
This time, we obtain a nice symmetric motif that matches the known FNR motif from RegulonDB. Click on the link [ alignments (logos): html on the right of the found motif. You can see the alignment as a logo with the known FNR motif.
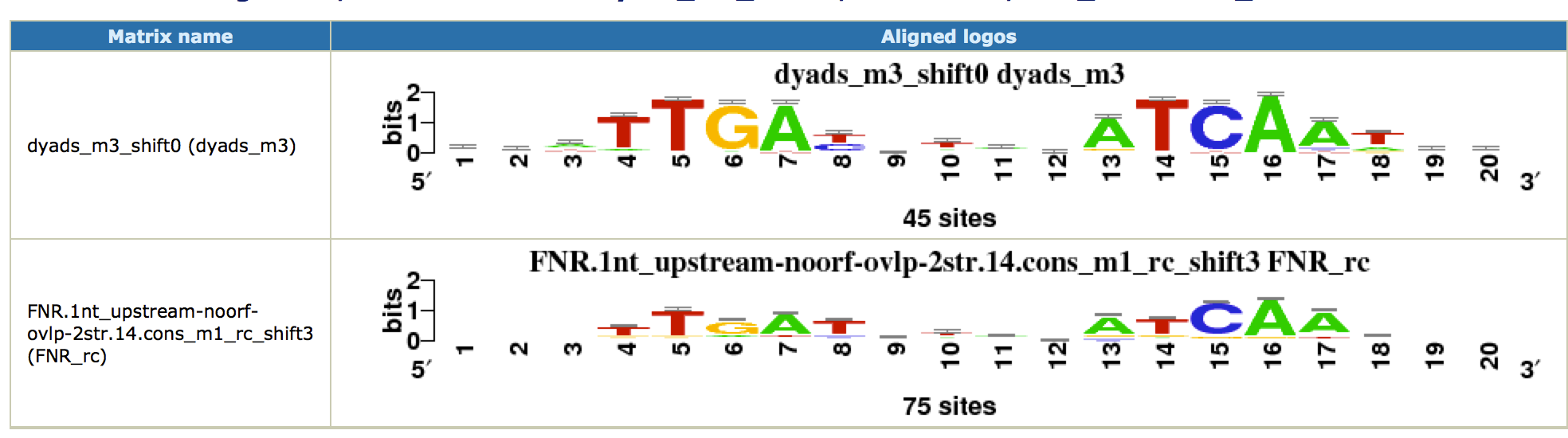

Note that this motif was only found by the dyad-analysis program, which highlights the importance of using several complementary algorithms for motif discovery. As there are no other factors, we did not find potential cofactors with this approach.
$ perl -lane '$start=$F[1]-100 ; $end = $F[2]+100 ; print "$F[0]\t$start\t$end"' macs14_summits.bed > macs14_summits+-100.bed
$ bedtools getfasta -fi Escherichia_coli_K_12_MG1655.fasta -bed macs14_summits+-100.bed -fo macs14_summits+-100.fa
FAQ
How to extract peaks from the supplementary data of a publication ?
The processed peaks (BED file) is sometimes available on the GEO website, or in supplementary data. Unfortunately, most of the time, the peak coordinates are embedded into supplementary tables and thus not usable "as is".
This is the case for the studied article. To be able to use these peaks (visualize them in a genome browser, compare them with the peaks found with another program, perform downstream analyses...), you will need to (re)-create a BED file from the information available.
Here, Table S5 provides the coordinates of the summit of the peaks. The coordinates are for the same assembly as we used.
- copy/paste the first column into a new file, and save it as retained_peaks.txt
- use a PERL command (or awk if you know this language) to create a BED-formatted file. As we need start and end coordinates, we will arbitrarily take +/-50bp around the summit.
- The BED file looks like this:
- Depending on the available information, the command will be different.
perl -lane 'print "gi|49175990|ref|NC_000913.2|\t".($F[0]-50)."\t".($F[0]+50)."\t" ' retained_peaks.txt > retained_peaks.bed
gi|49175990|ref|NC_000913.2| 120 220 gi|49175990|ref|NC_000913.2| 20536 20636 gi|49175990|ref|NC_000913.2| 29565 29665 gi|49175990|ref|NC_000913.2| 34015 34115
How to obtain the annotation (=Gene) BED file for IGV?
Annotation files can be found on genome websites, NCBI FTP server, Ensembl, ... However, IGV required GFF format, or BED format, which are often not directly available.
Here, I downloaded the annotation from the UCSC Table browser as "Escherichia_coli_K_12_MG1655.annotation.bed". Then, I changed the "chr" to the name of our genome with the following PERL command:
perl -pe 's/^chr/gi\|49175990\|ref\|NC_000913.2\|/' Escherichia_coli_K_12_MG1655.annotation.bed > Escherichia_coli_K_12_MG1655.annotation.fixed.bedThis file will work directly in IGV